The science
Ultupharma conducts research in two directions for antibiotics discovery and development:
1) Finding new antibacterial compounds in nature and optimizing their properties.
2) Developing new antibiotics by repurposing nucleoside analogues in combination with other known compounds.
----------------------------------
1) New antibacterial compounds in nature
Already Pasteur remarked that “la vie empêche la vie” but it was not until Fleming observed the chemical war between microorganisms that this resulted in the first potent antibiotic, penicillin.
Ultupharma has the goal to identify antibiotics with new mechanisms of action using microorganisms as sources to new metabolites. New microorganisms and new methods of cultivation developed in collaboration with a research group at SLU, have been found to produce new metabolites with anti-bacterial and anti-fungal effects. Initial screening for effect against bacteria resistant to present antibiotics and methods to rapidly determine chemical structures, novelty and patentability have been developed. More than 2000 microorganisms have so far been studied, allowing many thousand fractions to be tested annually. Several interesting new compounds have been identified with activities against bacteria and fungi causing disease in humans and animals.
A novel method for isolation of bacteria producing antibacterial peptides was recently published in Frontiers in Microbiology: Selective isolation of multidrug-resistant Pedobacter spp., producers of novel antibacterial peptides. This method resulted in the peptides of the ULT3-programme (ACS Chemical Biology: Isopedopeptins A–H: Cationic Cyclic Lipodepsipeptides from Pedobacter cryoconitis UP508 Targeting WHO Top-Priority Carbapenem-Resistant Bacteria). In the future, this approach is expected to yield many new antibiotic compounds for evaluation as putative new antibiotic drugs.
The ULT1 and ULT4-programmes are also based on compounds produced by microorganisms. The ULT1-compounds are described in Antibacterial 3,6-disubstituted 4-hydroxy-5,6-dihydro-2H-pyran-2-ones from Serratia plymuthica MF371-2, published in the Journal of Natural Products. The ULT1-compounds include two compounds isolated from the soil bacterium Serratia plymuthica, as well as a number of synthetic analogues. The ULT1-compounds are active against Gram-positive bacteria, including MRSA.
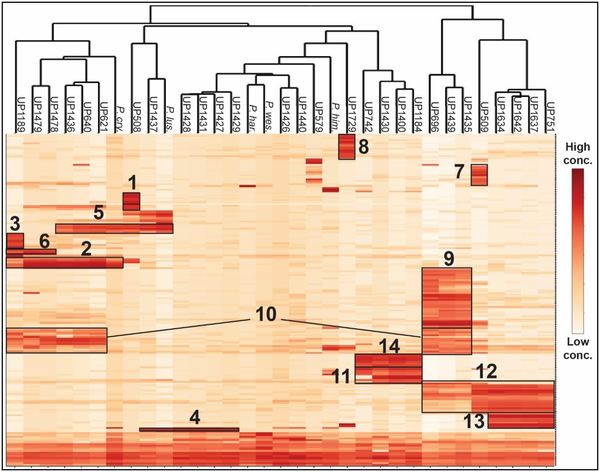
Left (click to enlarge): Hierarchical cluster analysis of antibacterial peptides produced by 28 Pedobacter sp. strains isolated from soil samples and five Pedobacter sp. type strains. Columns are bacterial strains and rows are peptides. Box 1 contains the ULT3 peptides and box 2-14 represent different types of peptides.
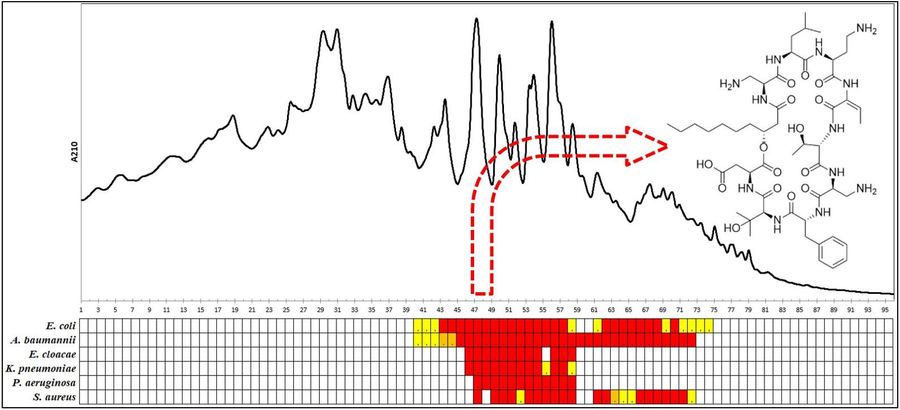
Right (click to enlarge): Bioassay-guided HPLC fractionation of culture extracts of Pedobacter cryoconitis UP508 yielding isopedopeptin B, one of the ULT3 peptides. Rows represent different test bacteria and columns are HPLC fractions.
Publications from the Ultupharma/SLU natural products screening efforts:
-Bjerketorp et al. 2021. Selective isolation of multidrug-resistant Pedobacter spp., producers of novel antibacterial peptides. Frontiers in Microbiology, 12, Article 642829.
-Broberg et al. 2021. In-peptide amino acid racemization via inter-residue oxazoline intermediates during acidic hydrolysis. Amino Acids 53(3), 323-331.
-Nord et al. 2020. Tetramic acid based alkaloids from Aspergillus amoenus Roberg strain UP197 – antibiotic properties and new pyranterreones. Natural Product Research, published on-line December 07, 2020.
-Nord et al. 2020. Isopedopeptins A–H: Cationic Cyclic Lipodepsipeptides from Pedobacter cryoconitis UP508 Targeting WHO Top-Priority Carbapenem-Resistant Bacteria. ACS Chemical Biology, 15, 2937-2944.
-Levenfors et al. 2020. Antibacterial pyrrolidinyl and piperidinyl substituted 2,4-diacetylphloroglucinols from Pseudomonas protegens UP46. The Journal of Antibiotics, 73, 739-747.
-Nord et al. 2019. Antibacterial isoquinoline alkaloids from the fungus Penicillium spathulatum Em19. Molecules 2019, 24, 4616.
-Bjerketorp et al. 2017. Antibacterial 3,6-disubstituted 4-hydroxy-5,6-dihydro-2H-pyran-2-ones from Serratia plymuthica MF371-2. Journal of Natural Products 80, 2997-3002.
-Broberg et al. 2017. Labradorins with antibacterial activity produced by Pseudomonas sp. Molecules, 22(7), 1072.
----------------------------------
2) Repurposing nucleoside analogues in combination with other known compounds
Repurposing nucleoside analogues, in combination with uridine and an inhibitor of bacterial dihydrofolate reductase, is the basis for the ULT2-programme.
The antibacterial profiles of the ULT2 combinations include broad activities against both Gram-negative and Gram-positive bacterial pathogens, thereby including many indications. The ULT2 programme has two additional key advantages; short time to market and low developmental risks. Several ULT2 combinations could be developed based on which indication(s) and bacteria(s) are targeted as described in the recent publication from Ultupharma Repurposing zidovudine and 5-fluoro-2′-deoxyuridine as antibiotic drugs made possible by synergy with both trimethoprim and the mitochondrial toxicity–reducing agent uridine in Journal of Antimicrobial Chemotherapy.
The mode of action is likely to involve several targets, including direct inhibition of the bacterial DNA polymerization and lowering the pool of thymidine available for DNA synthesis via several different mechanisms (Figure, below). The uridine component increases the effect of the other two components. In addition, it also decreases toxicity of the nucleoside analogues and has an anti-inflammatory effect and might reduce biofilm formation. This could be important in many infections where an anti-inflammatory effect can be beneficial. This includes sepsis, urinary tract infections, diabetic foot infections, Helicobacter pylori infections and several pulmonary infections. Uridine supplementation has been shown to have an anti-inflammatory and anti-fibrotic effect in an animal model of pulmonary fibrosis, and to improve intestinal barrier integrity and reduce incidence of diarrhea in piglets [Wu et al., J. Sci. Food Agric. 100:3709-3718 (2020)].
ULT2A and ULT2C are potent inhibitors of multiresistent E. coli, K. pneumoniae and S. aureus. The effect of both ULT2A and ULT2C against H. pylori offers new possibilities for cancer prophylaxis.

Figure. Suggested mode of action for combinations of azidothymidine (AZT) or 5-fluoro-2′-deoxyuridine (5-FdUrd) with trimethoprim (TMP) and uridine (U).
1) AZT reduces by competition the 2′-deoxythymidine-5′-monophosphate (dTMP) available for DNA synthesis.
2) AZT is phosphorylated to the DNA polymerase inhibitor AZT-5′-triphosphate (AZTTP).
3) TMP inhibits the modification of dihydrofolic acid (DHF) to tetrahydrofolic acid (THF). THF is transformed into 5,10-methylenetetrahydrofolate (5,10-CH2-THF), which is used for the formation of dTMP from 2′-deoxyuridine-5′-monophosphate (dUMP).
4) U inhibits the synthesis of 2′-deoxythymidine (dT) from thymine (Thy).
5) 5-FdUrd is phosphorylated to 5-FdUrd-5′-monophosphate (5-FdUrdMP), which is an inhibitor of dUMP methylation to dTMP.
6) U may via competition reduce the bacterial uptake of dT.
Modified from Levenfors et al, Repurposing zidovudine and 5-fluoro-2′-deoxyuridine as antibiotic drugs made possible by synergy with both trimethoprim and the mitochondrial toxicity–reducing agent uridine. Journal of Antimicrobial Chemotherapy, 2024, dkae438, https://doi.org/10.1093/jac/dkae438.
